For a detailed protocol, please refer to Nguyen et. al., JoVE 181, e63095 (2022)
Need guidance to implement iTACS on your microscope or looking for collaboration?
We will be glad to hear from you: dtambe@southalabama.edu
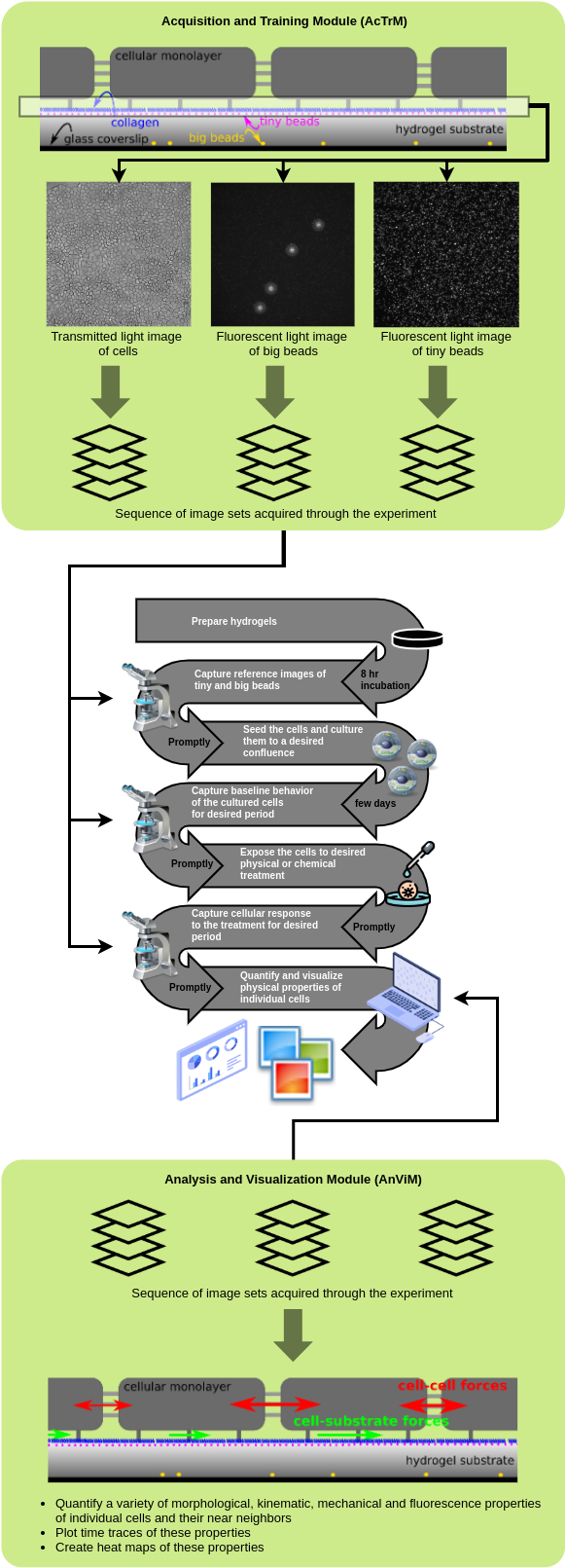
Here is an overview of an iTACS-based experiment to examine physiological properties of individual adherent cells:
Current version is developed and tested on Ubuntu operating system (18.04 - 20.10). One of the best ways to use the software on other operating system is through virtual Ubuntu machine which involves installing Oracle VM VirtualBox and on it install Ubuntu.
Software to compute cell-ECM forces
- Process used to compute cell-ECM forces is not included in this package, but any software that takes in the hydrogel surface deformation as input and provides forces exerted by a sheet of cells as output will work. Here is a software that will work for a thick hydrogel: FTTC.
- The cell-ECM forces are then used by the
inti,edge, andstripsoftware to quantify mechanical stresses within the cellular cluster. These software expect the cell-ECM forces to be inanalysis/p*/traction/*_trac.dat(where the first*is position number [0,1,2,3, ...] and the second*is the file number [0001,0002,0003, ...]) files with four columns:xyTxTy. Unit ofxandy: pixel andTxandTy: Pascal.
AcTrM is developed and tested on Micro-Manager-2.0beta on Windows 10.
| Task | Links |
|---|---|
| Install Micro-Manager 2.0 beta | https://valelab4.ucsf.edu/~MM/nightlyBuilds/2.0.0-beta/Windows |
| Establish a connection between Micro-Manager software and the microscope hardware | https://micro-manager.org/Device_Support https://micro-manager.org/Micro-Manager_Configuration_Guide |
Save a copy of Turboreg_.jar in the plugins folder of Micro-Manager |
http://bigwww.epfl.ch/thevenaz/turboreg/ |
In the plugins folder of Micro-Manager, create a subfolder IMBL and copy in this subfolder the files AcTrM.bsh, AcTrM_functions.bsh, and ColumnComparator.java |
https://github.com/IntegrativeMechanobiologyLaboratory/iTACS/blob/main/AcTrM/Micro-Manager-2.0beta/plugins/IMBL/AcTrM.bsh https://github.com/IntegrativeMechanobiologyLaboratory/iTACS/blob/main/AcTrM/Micro-Manager-2.0beta/plugins/IMBL/AcTrM_functions.bsh https://github.com/IntegrativeMechanobiologyLaboratory/iTACS/blob/main/AcTrM/Micro-Manager-2.0beta/plugins/IMBL/ColumnComparator.java |
- Create a folder
tnimgsin a suitable directory. Ensure that the path of this directory does not have space in the names of any of the parent folders. - Copy all the
c*_p*_t*.tiffiles from GitHub into thetnimgsfolder. - From Fiji, use
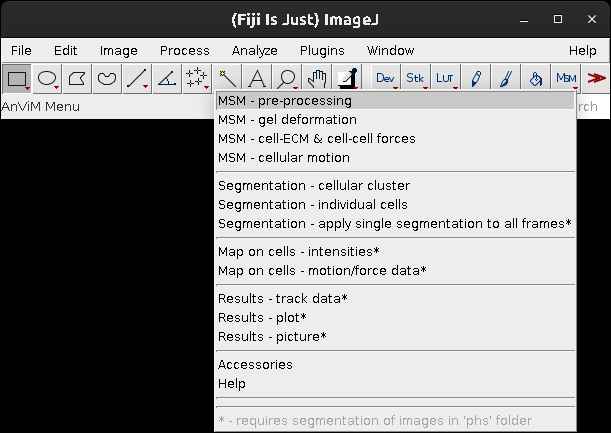
MSM>MSM - pre-processingto start data analysis and follow the procedure described in the Nguyen, Battle, Paudel et al., manuscript.
Nguyen, A., Battle, K.C., Paudel S.S., et al. Integrative Toolkit to Analyze Cellular Signals: Forces, Motion, Morphology, and Fluorescence. J. Vis. Exp. 181, e63095 (2022).
PDF
HTML
To segment smooth muscle cells that are separated and have long and narrow extensions,
- conduct cluster segmentation by thresholding to have white cells (pixel value 255) and black background (pixel value 0).
- for individual segmentation, start with parameter values: sledgehammer = 0, proninance = 100, mallet = 40